Scientists find new system in tomato’s defense against bacterial speck disease
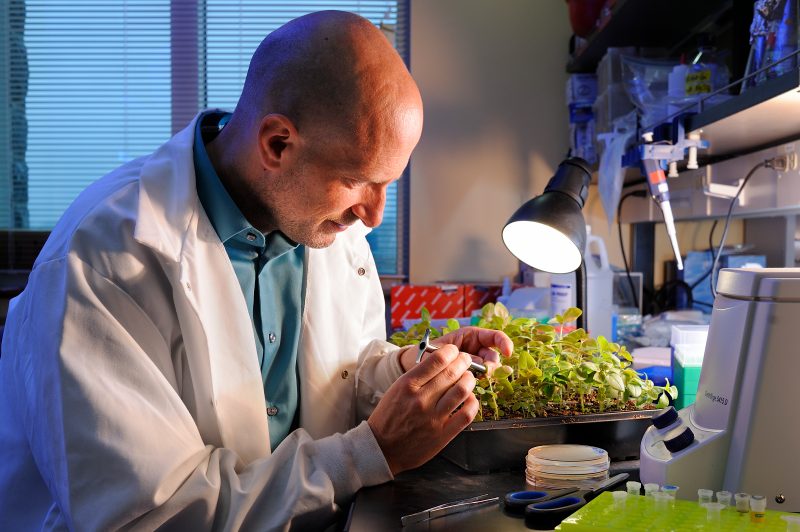

Virginia Tech researchers, along with the Boyce Thompson Institute, have discovered a new mechanism in the continual arms race between plants and pathogenic bacteria. The team identified a new receptor in tomato plants, called FLAGELLIN-SENSING 3 (FLS3), that triggers defenses against a bacterial attack. The study appears in Nature Plants.
The discovery adds to a growing body of knowledge on how plants — as well as animals and humans — evolved to efficiently defend themselves against bacterial pathogens by detecting conserved bacterial molecules, such as the building block of the bacterial flagellum, called flagellin.
“This discovery sets up the possibility of introducing FLS3 into other economically important crop plants, which might provide resistance to bacterial pathogens that is not naturally present in these other plants” said first author Sarah Hind, a research associate in the lab of Professor Gregory Martin at BTI, which is located on the Cornell University campus.
Boris Vinatzer, a professor of plant pathology, physiology, and weed science in the Virginia Tech College of Agriculture and Life Sciences, and Christopher Clark, a post-doctoral researcher in Vinatzer’s lab, collaborated on the project.
Vinatzer’s laboratory previously identified flgII-28, the conserved region of the bacterial flagellin molecule that FLS3 recognizes, by studying the bacterial speck pathogen.
“This discovery shows how the study of the genetic diversity of a pathogen can lead to a deeper understanding of the plant’s immune system, which in turn could be used to develop disease-resistant crops,” Vinatzer said.
FLS3 is the second flagellin sensor discovered in tomatoes. The first, called FLS2, is found in most land plants and detects invading bacteria through recognition of a separate region of flagellin called flg22. Since many plants have an FLS2 receptor, several bacterial species have acquired mutations in the flg22 portion of their flagellin, and these mutations alter the flagellin shape enough that FLS2 can no longer recognize it. The acquisition of the FLS3 receptor by the ancestor of the modern tomato may therefore serves as a countermeasure on behalf of the tomato to detect bacteria that have altered flg22.
The study demonstrates how versatile the plant immune system can be while fighting a constant arms race against infectious bacteria.
“Plants are always coming up with new ways to defeat pathogens,” said Hind. “We’re trying to understand how they do it and then use this knowledge to develop more disease-resistant plants.”
Research reported in this news release was supported by the National Science Foundation, the USDA National Initiative in Food and Agriculture, the USDA Binational Agriculture Development Fund, the National Institutes of Health, the TRIAD Foundation, the Human Frontiers Science Program, and through internal funding from the Boyce Thompson Institute.