Tuberculosis infection in mongoose driven by social communication behavior
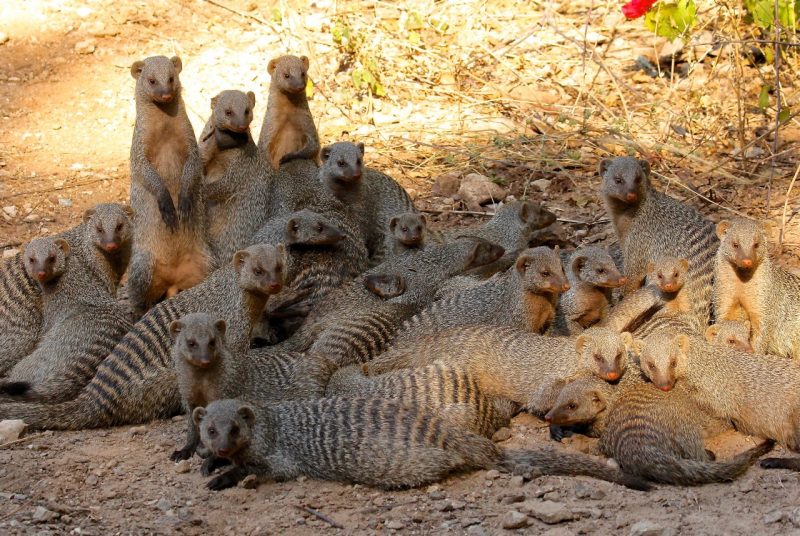

An emerging strain of tuberculosis (TB), closely related to human TB, has been killing banded mongoose in Northern Botswana in significant numbers. This novel pathogen, Mycobacterium mungi, did not infect mongoose through a primary airborne or oral route as normally seen in TB disease in humans and animals. The mechanism of transmission, however, was unknown.
Now, a research team led by Kathleen Alexander, associate professor of wildlife conservation in Virginia Tech’s College of Natural Resources and Environment, reports discovery of the pathogen’s unique transmission route in the May 10 issue of the American Society for Microbiology journal mBio.
Using a suite of molecular techniques to identify the presence of M. mungi-specific DNA and examination of mongoose tissues and cells, Alexander and her team have discovered that TB transmission in mongoose occurs in conjunction with social behavior. As with many animals, such as dogs or even hyenas, mongoose use urine and anal gland secretions to communicate with other members of their species. However, in the mongoose, secretions from sick animals were found to be infected with the TB pathogen.
These secretions, once deposited in the environment, allow the pathogen to be transmitted when other mongoose investigate and sniff the scent marks. The pathogen is also spread when an infected mongoose places its scent directly on other mongoose in its troop. Abrasions or injuries in the skin or nose provide the portal of entry for this novel TB pathogen to invade and infect the mongoose host. Smaller social groups are most threatened by the disease, the researchers report.
“Banded mongoose are a territorial species, and individuals within a troop may have little or no direct contact with mongoose in adjacent social groups, limiting the potential for directly transmitted pathogens like TB to spread through a population,” explained Alexander, an affiliate of the Fralin Life Science Institute, who discovered the novel strain of TB in 2010.
“But this TB pathogen circumvents the mongoose’s natural social barriers to infectious disease transmission by hijacking social communication behavior,” she said. “We keep being surprised by infectious disease-causing organisms and their ability to adapt to a particular environment, behaving, in some cases, dramatically differently than we expect.”
TB is an ancient disease that continues to be one of the most important health threats to humans, wildlife, and domestic animals globally. The discovery by Alexander’s team of the novel mode of infection by M. mungi in banded mongoose has critical implications to the current understanding of tuberculosis infection dynamics, warranting further examination of other species where this transmission pathway may also occur, the researchers point out in their article.
Potential sources of pathogen exposure were evaluated, including soil, sewage, and human and mongoose feces, as well as feces from 16 different wildlife species — from elephants to domestic cows. Despite this, M. mungi DNA could only be found in banded mongoose tissues and secretions. The scientists examined 155 mongoose between July 2000 and June 2015, conducting in-depth studies of tissues from 79 of these animals.
TB lesions were found in a variety of organs, but more significantly in the nose, nasal cavity, and skin — parts of the mongoose host that make frequent contact with anal gland secretions and urine during olfactory communication behavior. Lung lesions were only found in affected animals in advanced stages of the disease.
“M. tuberculosis complex pathogens infect many species of domestic and wild animals as well as humans in the U.S. and across the globe,” Alexander said. “Our findings have changed the way we must think about tuberculosis and infectious disease transmission in territorial species.”
“Mechanisms of host exposure are still not completely understood for many host species and M. tuberculosis complex organisms,” she continued. “There is an urgent need to better understand the processes that influence environmental transmission and persistence of TB pathogens and resultant disease control implications.”
Alexander noted, “We have recently sequenced the genome of this emerging pathogen, and we can now start to investigate why this TB pathogen behaves so differently – patterns that have important implications to our understanding of TB disease in both humans and animals.”
Members of the research team also include Claire Sanderson, research associate in fish and wildlife conservation at Virginia Tech; Michelle Larsen of the Department of Medicine, Albert Einstein College of Medicine; Suelee Robbe-Austerman of the Diagnostic Bacteriology Laboratory, National Veterinary Services Laboratories, Ames, Iowa; Mark C. Williams of the University of Pretoria, Onderstepoort, South Africa; and Mitchell Palmer of the Bacterial Diseases of Livestock Research Unit, National Animal Disease Center, Ames, Iowa.
The work was supported by a Morris Animal Foundation grant and a National Science Foundation grant as part of the joint National Science Foundation–National Institutes of Health–U.S. Department of Agriculture Ecology and Evolution of Infectious Diseases program.
Alexander, a wildlife veterinarian and co-founder of the Center for Conservation of African Resources: Animals, Communities, and Land Use (CARACAL) in Botswana, directs her research program at exploring and understanding the factors that influence the emergence and persistence of novel and re-emerging diseases at the human-wildlife-environment interface.