Virginia Tech researchers discover insights into HIV with implications for drug development

Though human immunodeficiency virus, or HIV, was discovered more than 35 years ago, and more than 36 million people around the world are living with the virus today, a vaccine still remains elusive.
But a Virginia Tech research team has discovered insights that can provide perspective on how to design drugs that can more effectively target the surface of the virus and ultimately block HIV from infecting cells.
“We hope this new insight on the druggable surface of HIV will provide routes to novel types of therapeutics to combat this virus that has taken so many lives,” said Anne M. Brown, an assistant professor in research and informatics in University Libraries and an adjunct professor in the Department of Biochemistry in the College of Agriculture and Life Sciences.
Her team’s research focuses on computer modeling and integrating these computational tools into biological and life science research. Their recent findings were published in Biophysical Journal.
Brown collaborates with many research groups on campus to integrate computational and informatics techniques into their research and teaching and is an affiliate of the Fralin Life Science Institute and the Virginia Tech Center for Drug Discovery.
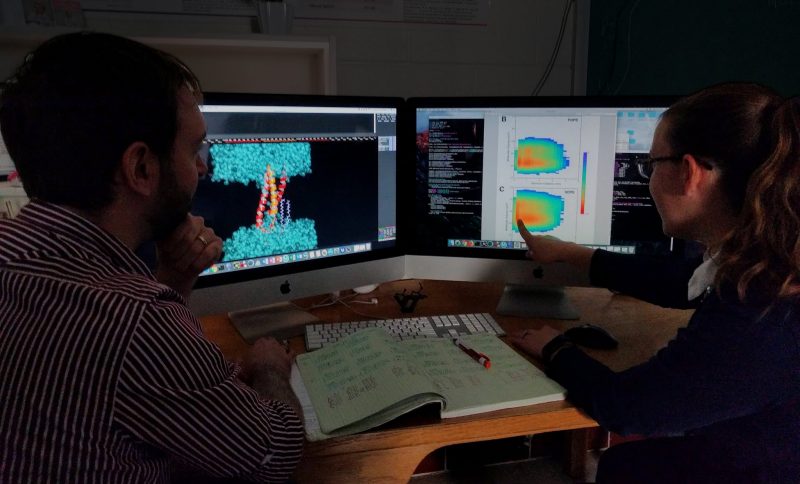
For this work, the team focused on the gp41 transmembrane domain (TMD) of the HIV surface protein, which reaches out like a spike to help the virus attach to and infect cells; the surface of the virus is the only viable target for anti-HIV vaccines. The group performed molecular dynamics simulations of TMD in a model asymmetric viral membrane that mimics the native membrane of HIV.
"This simulation of the TMD is particularly significant because it allows us to observe structural changes that occur in the TMD that can then be transmitted to other parts of the surface protein. A detailed understanding of these changes is critical to rational design of therapeutics directed against HIV," said David Bevan, a professor of biochemistry in the College of Agriculture and Life Sciences.
Brown completed her Ph.D. at Virginia Tech in Bevan’s lab and now runs a collaborative lab with Bevan. As of the spring 2018 semester, they had 25 undergraduate students working in their lab.
Louis “Bobby” Hollingsworth, a former undergraduate researcher in the lab and first author on the paper, said looking at the virus on an atomic scale gives new insight into how the TMD region of the virus might interact and respond to possible drug treatments.
“This atomistic picture of TMD adds to the body of knowledge on HIV biophysics and mechanisms, hopefully contributing to further studies and therapies,” said Hollingsworth, who is currently a graduate student at Harvard University and Boston Children’s Hospital.
The team found that water and chloride ions permeated the membrane and interacted with the highly conserved arginine bundle (R696)3 at the center of the membrane and influenced TMD stability. If researchers can destabilize these areas of TMD with drug design, they can ultimately block HIV infection.

Next, Brown’s team will complete more models of TMD and its interaction with potential drugs that may block viral entry into human cells. They can then collaborate with immunologists to confirm these models in a wet lab environment and test the potential therapeutics. They also seek to use these methods to better understand other transmembrane proteins.
Justin Lemkul, an assistant professor of biochemistry in the College of Agriculture and Life Sciences, was a contributing author on this paper and coded and scripted some of the membrane simulations for this project.
“The more we can understand about how the proteins in HIV move, change shape, and interact with viral and cellular membranes, the better chance we have at developing therapies against the virus,” he said.



.jpg.transform/m-medium/image.jpg)

